Network Degeneration
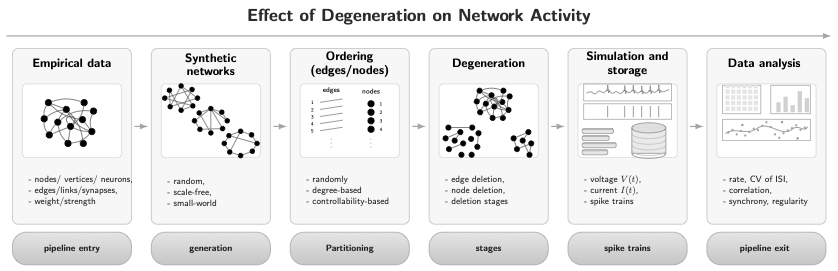
Reproducible pipeline to study how staged structural degeneration alters dynamical activity in spiking neuronal networks.
Details · Repository — Private until published; available upon request
Problem
Quantify how controlled network degradation (synapse pruning and neuron deletion) changes spiking dynamics and structure-function observables across parameter grids.
Approach
The pipeline is structured in two phases. In presimulation, it generates or loads directed E/I networks, applies a deterministic edge ordering (via maximum matching decomposition), performs staged degeneration (synaptic pruning and neuron deletion), and assigns block-wise weights (EE, EI, IE, II scaling) to produce weighted connectivity instances.
In simulation and postsimulation, each fixed network instance is simulated (currently via NEST), producing spike trains and time series. The pipeline then extracts activity-based and structure-based observables and stores them as structured artifacts for aggregation across parameter grids.
Impact
Demonstrates system design for computational experiments: deterministic preprocessing, scalable simulation runs, and structured outputs enabling downstream statistical analysis.
Key outputs
- Degenerating network instances (edges/nodes/weights)
- Spike trains and dynamical time-series from NEST simulations
- Activity metrics: firing rates, ISI CV, synchrony, pairwise correlations
- Structure metrics: degree statistics, weighted sums, common inputs, spectral radius, block summaries
- Structured .npz artifacts aggregated over parameter grids
Tools and Methods
- Python
- Graph algorithms
- NumPy / NPZ datasets
- Parallel execution
Planned extensions
- Formal workflow orchestration (Snakemake)
- Feature correlation and prediction analysis